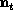 at time
t. The age-stage distribution evolves according to
at time
t. The age-stage distribution evolves according to
Session III: Population Dynamics of Target Species
Charlie Miller, Chair
Cabell Davis & Dan Lynch, Rapporteurs
Development of Stage-Structured Models of Fish Community Dynamics
Michael J. Fogarty
National Marine Fisheries Service, Woods Hole
The development of models for multispecies fisheries systems has been dominated by two general approaches: (1) Lotka-Volterra type models with modifications to account for removals due to harvesting and (2) detailed age-structured models with explicit consideration of harvesting and interspecific interactions. The former cannot accommodate differences among life history stages or size classes within populations and communities while the latter require extensive information on age composition for all members of the community. Stage- structured models provide an alternative approach which preserve important features related to life stages or size classes. Critical demographic and ecological factors are often more closely related to ontogenetic stage or size than to age.
Two stage-structured models under development for describing fish community dynamics on Georges Bank funded by the NOAA Coastal Ocean Program are illustrated. The first is a simple two-stage model comprising juveniles and adults of a group of interacting species. This model is defined in discrete time in a delay-difference formulation. The model describes survivorship from the previous time step and the addition to the population through recruitment. Interspecific interactions can occur at both the juvenile and adult stages. For highly nonlinear recruitment functions, a broad array of dynamical behavior including stable points, cycles and chaos is possible for even simple two species systems. Harvesting and interspecific interactions are shown to have synergistic effects on the dynamics of the system. In particular, harvesting juveniles reduces the overall resilience (persistence of all species) of the system but also reduces the probability of observing complex dynamical behavior. An illustration of applying this model to predator-prey interactions in a two species system (cod-herring) is provided.
The second model is an extension of the first to include multiple life history stages and/or size classes and spatial structure. This formulation provides greater flexibility in describing the population and community dynamics for the case where nonlinear processes are important at several points in the life cycle and/or where spatial aggregates with little intermixing among groups are known to occur. Such models can readily give rise to multiple stable states in community structure. These models are also considerably
more demanding with respect to data requirements. The objective of NOAA Coastal Ocean Program studies on Georges Bank is to develop models of exploited multispecies systems in support of fishery management which can accommodate these factors. A combination of process-oriented field and laboratory, and is underway to define and estimate critical parameters for these models.
Oligonucleotide-Probe Based Identification of Sibling Species and their Application for Population Dynamic Studies
Ann Bucklin1, Eva P. Curry1, and Nancy J. Copley2
1University of New Hampshire
2Woods Hole Oceanographic Institution
One of the goals of the Georges Bank Study is to understand and uantitatively describe the population ecology of the copepods, Peudocalanus moultoni and P. newmani, that together play an important role in the Bank ecosystem. In order to fully understand the ecosystem dynamics, it is essential to differentiate these two very similar, sympatric marine species and to work toward understanding the processes that keep the species distinct. Discrimination of Pseudocalanus species will enable estimation of their respective secondary production, and will allow understanding of the biological and physical determinants of their distribution and abundance on Georges Bank.
Although P. moultoni and P. newmani are extremely similar in morphological characters and in many aspects of their ecology, they apparently maintain reproductive isolation even in areas of sympatry (Hart and McLaren, 1978; Frost, 1989; Sevigny et al., 1989). They are markedly distinct in genetic character, based on the DNA sequence of portions of themitochondrial genes, 16S rRNA and cytochrome oxidase I (COI) (Bucklin et al., 1995). Oligonucleotide probes have been designed from regions of COI that are at least 30% different from all other analyzed copepods and are currently be tested for specificity in dot-blot hybridization reactions. These probes should discriminate Pseudocalanus spp. larvae and juveniles not only from all other species but also from each other. Since the first step in the hybridization process is amplification by polymerase chain
reaction (PCR), individual eggs, larvae, and juveniles can be used for molecular analyses. Hundreds of reactions can be hybridized simultaneously; the entire hybridization process requires approximately 2 days' intermittent effort.
Oligonucleotide probes can be used to detemine the seasonal timing, hydrographic conditions, and geographic location of reproduction of each Pseudocalanus species. The temporal and spatial patterns of reproduction of P. newmani and P. moultoni can then be determined from the distribution and abundance of larval and juvenile stages, in comparison to adult distributions.
References:
Bucklin, A., E. Curry, and N.J. Copley (1995) Molecular phylogeny and species identification of planktonic calanoid copepods of Georges Bank based on mitochondrial 16S rRNA gene sequences. Mar. Biol. (in press)
Frost, B.W. (1989) A taxonomy of the marine calanoic copepod genus Pseudocalanus. Can. J. Zool. 67:525-551.
Hart, R.C. and I.A. McLaren (1978) Temperature acclimation and other influences on embryonic duration in the copepod Pseudocalanus sp. Mar. Biol. 45:23-30.
Sevigny, J.-M., I.A. McLaren, and B.W. Frost (1989) Discrimination among and variation within species of Pseudocalanus based on the GPI locus. Mar. Biol. 102:321-327.
Larval cod (Gadhus morhua) and haddock (Melannogrammus aeglefinus) growth on Georges Bank: a model with temperature, prey size and turbulence forcing
Andrew W. Leising and Peter J.S. Franks
Scripps Institution of Oceanography
Previous individual-based modeling studies have investigated the effects of variability in prey density and turbulence on first feeding cod and haddock larvae (Werner et al., 1994). We have developed an individual-based model, based on the model of Laurence (1985), that includes these factors and the effect of varying temperature. Temperature effects were included by using a Q10 type adjustment to the metabolic rate. Besides affecting metabolic weight loss, temperature also may affect the swimming speed, capture efficiency, and general "liveliness" of a fish (Brown et al., 1989). Therefore, a second temperature dependent term was added to the overall ingestion ability of our model fish. We have also developed a scheme to convert total prey mass per unit volume to a discrete number of prey items for the larval fish to encounter. Three cases were analyzed: 1) constant food and temperature conditions, 2) variable temperature cycles, and 3) variable temperature cycles and turbulence. Our model results showed that both the timing and location of cod and haddock spawning are crucial to the survival of the larvae (Figure 1). We also found that it is important to force the model with the temperature cycle and not just the average temperature over that cycle (Figure 2). Finally, turbulence may increase growth rates in clod locations where growth is limited by food and temperature, by increasing encounters with prey (Figure 2).
Figure 1: Locations on Georges bank where larvae were modeled. (Figure modified from Laurence and Lough, 1985).
Figure 2: Growth of survivors of 100 cod larvae run for 40 days.
Reduction of complexity in biological/physical models
Glenn R. Flierl, MIT, and Cabell S. Davis, WHOI
Biological models of the life-cycle of an organism or of many interacting species are often complex with large numbers of variables, and, therefore, cannot be incorporated into a full two- or three-dimensional model of a oceanic region. This paper presents a technique for reducing the complexity of such a model (using as an example, the model of Davis, 1984, for the life cycle of the copepod Pseudocalanus), while retaining the essential aspects. The full model is run in a system with no spatial variation, but with forcing representing the expected influence and time scales of the physical forcing. From this calculation, the Empirical Orthogonal Functions (EOF's), representing the principal temporal and age-stage variability, are extracted and a small set of the age-stage EOF's are used as basis functions. Simplified dynamics are derived by projecting the original dynamical system onto this set of basis functions.
Mathematically, the procedure can be expressed using an example:
suppose we begin with a zero-dimensional model for the age-stage
structure (represented as a vector 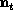 at time
t. The age-stage distribution evolves according to
at time
t. The age-stage distribution evolves according to
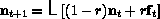
where r is the exchange rate with the
exterior and 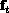 is the external population
(specified). From this, we build up a matrix
is the external population
(specified). From this, we build up a matrix  of
age-stage X time. To represent this with EOF's, we find two matrices
of
age-stage X time. To represent this with EOF's, we find two matrices
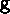 (age-stage X EOF number) and
(age-stage X EOF number) and 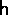 (EOF number X time) such that the product
(EOF number X time) such that the product 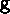
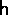 optimally fits
optimally fits  . We then use the
vectors in
. We then use the
vectors in 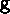 as a new basis function and project the
dynamics, approximating
as a new basis function and project the
dynamics, approximating 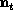 by
by 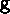
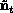 where the order of
where the order of 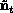 is the number
of EOF's retained, rather than the much larger number of age-stage
classes. The EOF amplitudes evolve according to
is the number
of EOF's retained, rather than the much larger number of age-stage
classes. The EOF amplitudes evolve according to
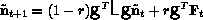
which is the same form as the original model, but reduced in dimension.
We examine development of an initial pulse, flow over one- and two-dimensional topography, and nonlinear interactions with a food source. In all these problems, the full 200 variable system can be adequately reproduced using between 5 and 15 modes (Figure 1). The reduced models are generally much more accurate than a "grouped" model (e.g., an Eggs- Nauplii- Copepodid- Adult representation). A second advantage is that the EOF reduction does not have adjustable parameters (other than the number of modes to include). The EOF approach allows the important aspects of detailed biological interactions (i.e., more realism) to be included in a large-scale physical model.
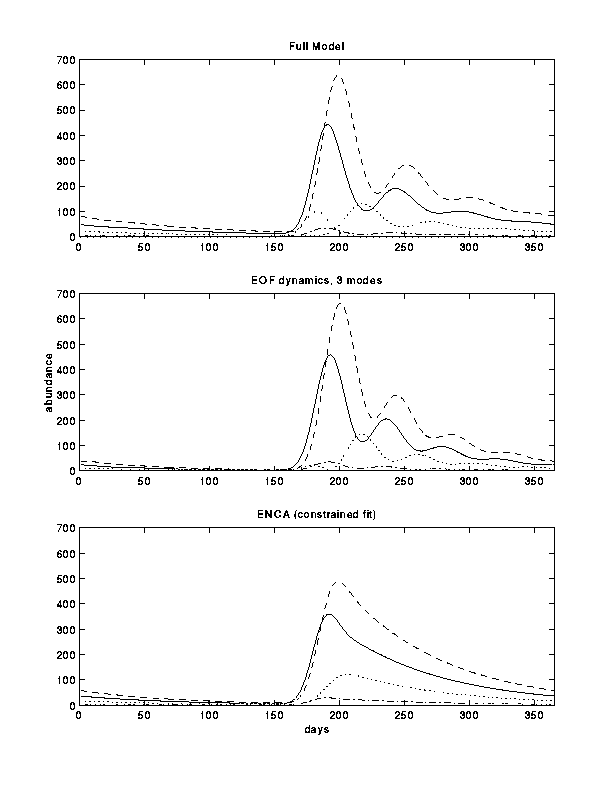
Figure 1: Population structure resulting from a yearly pulse in the external population calculated with various models. The lines show the number of Eggs (solid), Nauplii (dash), Copepodids (dot) and Adult females (dash-dot). The heavy dots show the external adult population versus time (multiplied by 100).